Report into a nosocomial outbreak of coronavirus disease 2019 (COVID-19) at Netcare St. Augustine's Hospital
This report presents the findings and recommendations of an investigation into a nosocomial outbreak of coronavirus disease 2019 (COVID-19) at St. Augustine's Hospital in Durban, South Africa. The investigation began on 4 April after the identification of a number of confirmed COVID-19 cases and three deaths at the hospital. Investigation methods included medical record reviews, ward visits, and interviews with health care workers and management. A detailed timeline of patient cases was constructed to generate hypotheses as to the spread of infection through the hospital. In addition, DNA sequencing of severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) nucleic acid extracted from nasopharyngeal and oropharyngeal swab samples was performed and phylogenetic analysis was conducted.
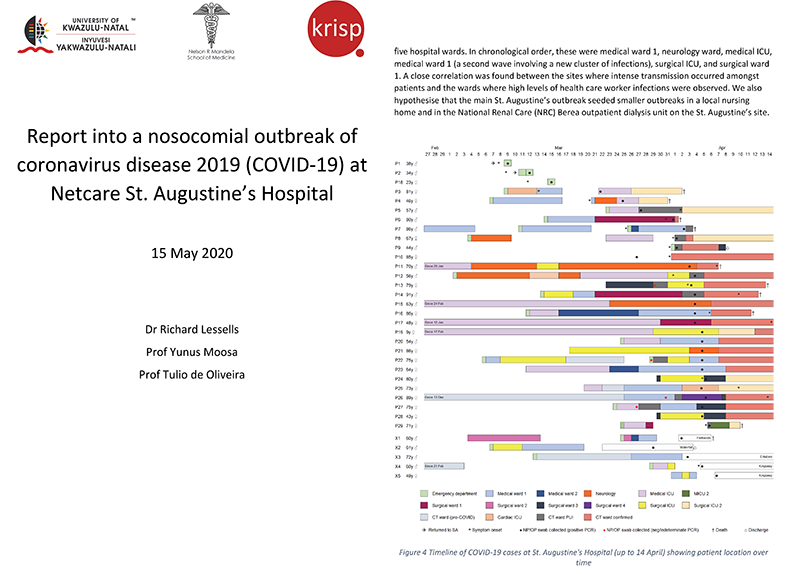
Between 9 March and 30 April 2020, 119 confirmed cases were identified at St. Augustine's Hospital (39 patients and 80 staff). The 80 staff represent approximately 5% of all staff tested for SARS-CoV-2. The most plausible hypothesis is that there was a single introduction of SARS-CoV-2 to the hospital on 9 March, most likely as a result of transmission from a patient attending the Emergency Department for investigation of COVID-19 to another patient present in the ED at the same time who was then admitted to the cardiac intensive care unit with a suspected stroke. The infection then spread rapidly through the hospital, involving patients on at least five wards.Phylogenetic analysis supports the main hypothesis of a unique introduction followed by widespread transmission in the hospital. All of the 18 SARS-CoV-2 genomes produced (nine from patients and nine from health care workers) clustered together with limited genetic diversity. All of the sequences belong to the A2a clade associated with infections from Europe.
In response to the coronavirus disease 2019 (COVID-19) pandemic, we created the Network for Genomics Surveillance in South Africa (NGS-SA) with a grant from the South African Medical Research Council (SAMRC) and the South African Department of Science and Innovation (DSI). Our goal is to sequence at least 10,000 genomes of severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) to inform the public health response in South Africa.
 >
>News date: 2020-05-22
Links:
KRISP has been created by the coordinated effort of the University of KwaZulu-Natal (UKZN), the Technology Innovation Agency (TIA) and the South African Medical Research Countil (SAMRC).
Location: K-RITH Tower Building
Nelson R Mandela School of Medicine, UKZN
719 Umbilo Road, Durban, South Africa.
Director: Prof. Tulio de Oliveira




