KRISP work to produce a bioinformatics software for coronavirus COVID-19 rapid identification and characterization
We developed an rapid bioinformatics tool for the identification and characterization of novel coronavirus genomes. The tool was released in January 2020 and published in Bioinformatics in February 2020 as an open access tool to help to characterise genomes of COVID-19 viruses.
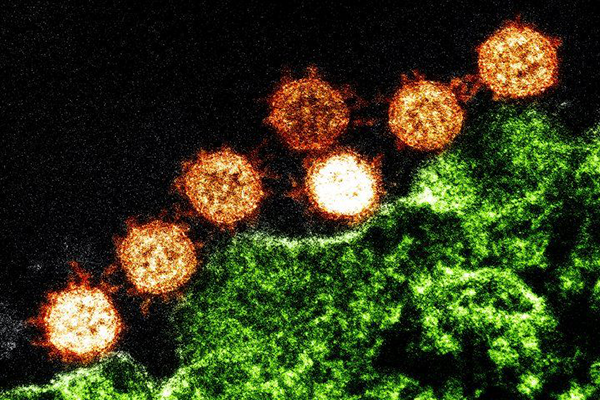
We have been following up on the COVID-19 outbreak since the first sequences for the SARS-CoV-2 virus have been made publicly available on the 11th of January 2020. We added some exciting new features to Genome Detective to support SARS-CoV-2 better.
On January 15th, the first test release of the Coronavirus typing tool was available. On the 31st of January, a full paper was deposited openly in biorxv. This paper received over 3,000 accessions and on the 17th of February was accepted for publication in the Bioinformatics journal - Sara Cleemput et al. Bioinformatics 2020.
All of our results and software were released in real-time and the Nature journalist Jack Leeming wrote an interesting article on Nature Index (February 26th) highlighting we developed the tool to to respond quickly to this coronavirus outbreak. This piece also hoghlith how our previous work on Yellow Fever Virus and Zika (published at Science and PLoS NTD) have prepared us
We have been improving Genome detective continuously, such as by improving mutation calling, and reassembly of raw viral Oxford Nanopore sequence data. UZLeuven used Genome Detective for confirming the diagnosis of the first Belgian SARS-CoV-2 infected patient, see use case in the video below.
News date: 2020-03-04
Links:
https://www.krisp.org.za/publications.php?pubid=272
Genome Detective Coronavirus Typing Tool for rapid identification and characterization of novel coronavirus genomes. Cleemput S, Dumon W, Fonseca V, Abdool Karim W, Giovanetti M, Alcantara LCJ, Deforche K, de Oliveira T, Bioinformatics (2020), btaa145, https://doi.org/10.1093/bioinformatics/btaa145:.
KRISP has been created by the coordinated effort of the University of KwaZulu-Natal (UKZN), the Technology Innovation Agency (TIA) and the South African Medical Research Countil (SAMRC).
Location: K-RITH Tower Building
Nelson R Mandela School of Medicine, UKZN
719 Umbilo Road, Durban, South Africa.
Director: Prof. Tulio de Oliveira





