Coronavirus variants get Greek names, but will scientists use them?
From Alpha to Omega, the labelling system aims to avoid confusion and stigmatization. To quell such confusion and avoid geographical stigmas, everyone should now just call it Beta according to a naming scheme announced on 31 May by the World Health Organization (WHO) in Geneva and described in a forthcoming article in Nature Microbiology.

Colourized scanning electron micrograph of a cell (red) infected with SARS-COV-2 virus particles (yellow).Credit: NIAID
When researchers in South Africa spotted a highly mutated strain of coronavirus driving the country’s second wave in late 2020, they called it variant 501Y.V2. Naming schemes developed by other scientists have called it B.1.351, 20H/501Y.V2 and GH/501Y.V2. But many media outlets — and some scientists — describe the same virus as ‘the South African variant’.
To quell such confusion and avoid geographical stigmas, everyone should now just call it ‘Beta’, according to a naming scheme announced on 31 May by the World Health Organization (WHO) in Geneva and described in a forthcoming article in Nature Microbiology.
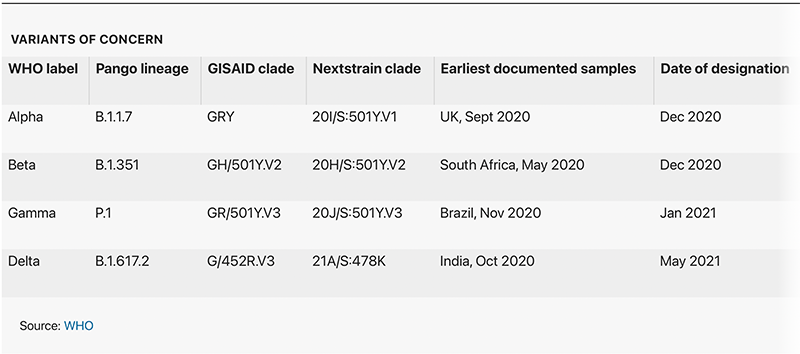
The names, taken from the Greek alphabet (see ‘Variants of concern’), are not intended to replace scientific labels, but will serve as a handy shorthand for policymakers, the public and other non-experts who are increasingly losing track of different variant names.
'It is a lot easier for a radio newsreader to say ‘Delta’ than bee-one-six-one-seven-two,' says Jeffrey Barrett, a statistical geneticist leading SARS-CoV-2-sequencing efforts at the Wellcome Sanger Institute in Hinxton, UK. 'So I’m willing to give it a try to help it take off.'
'LetÂ’s hope it sticks,' says Tulio de Oliveira, a bioinformatician and director of the KwaZulu-Natal Research Innovation and Sequencing Platform in Durban, South Africa, whose team identified the Beta variant. 'I find the names quite simple and easy.'
The system could be especially useful in countries battling a number of variants, such as South Africa, where a variant found in the United Kingdom and known to scientists as B.1.1.7 — now called Alpha — is on the rise, and researchers such as de Oliveira are watching out for cases of the B.1.617.2 variant identified in India, now called Delta. 'For a country like South Africa, to follow Beta and Alpha and keep a small eye on Delta, that will potentially be easier,' he says.

Confusion isn’t the only reason to go with a simplified naming system, say advocates of the new system. Terms such as ‘the South African variant’ and ‘the Indian variant’ can stigmatize countries and their residents, and might even discourage nations from running surveillance for new variants. 'The geographical names, we have to stop with that — really,' says de Oliveira. He is aware of countries in Africa where health ministers have been reluctant to announce the discovery of new local variants because of concerns about being made pariahs.
'I can understand why people just call it ‘the South African variant’ — they don’t mean anything by it,' says Salim Abdool Karim, an epidemiologist at the Centre for the AIDS Program of Research in South Africa in Durban. 'The problem is, if we allow it to continue, there are people who have an agenda and will use it.'
Barrett intends to embrace the new naming system in media appearances, but he suspects geographical descriptors wonÂ’t go away quickly. 'The reason we use country names (which is problematic) is that it ties the variants to the story of the pandemic in a way thatÂ’s easier to remember,' he wrote in an e-mail to Nature. 'The new system is still very anonymous and it will still be hard for the public to remember whoÂ’s who.'
In recent months, most scientists have settled on a single lineage-naming system that describes the evolutionary relationships between variants. With time, the WHOÂ’s naming system might gain the same currency among the general public, says Jeremy Kamil, a virologist at Louisiana State University Health in Shreveport. 'If people use it, it will become the default.'
Nature 594, 162 (2021)
doi: https://doi.org/10.1038/d41586-021-01483-0
News date: 2021-06-01
Links:
https://www.nature.com/articles/d41586-021-01483-0
SARS-CoV-2 Variants of Interest and Concern naming scheme conducive for global discourse. Konings F, Perkins MD, Kuhn JH, Pallen MJ, Alm EJ, Archer BN, Barakat A, Bedford T, Bhiman JN, Caly L, Carter LL, Cullinane A, de Oliveira T, Druce J, El Masry I, Evans R, Gao GF, Gorbalenya AE, Hamblion E, Herring BL, Hodcroft E, Holmes EC, Kakkar M, Khare S, Koopmans MPG, Korber B, Leite J, MacCannell D, Marklewitz M, Maurer-Stroh S, Rico JAM, Munster VJ, Neher R, Munnink BO, Pavlin BI, Peiris M, Poon L, Pybus O, Rambaut A, Resende P, Subissi L, Thiel V, Tong S, van der Werf S, von Gottberg A, Ziebuhr J, Van Kerkhove MD, Nature Microbiology (2021), https://doi.org/10.1038/s41564-021-00932-w:.
KRISP has been created by the coordinated effort of the University of KwaZulu-Natal (UKZN), the Technology Innovation Agency (TIA) and the South African Medical Research Countil (SAMRC).
Location: K-RITH Tower Building
Nelson R Mandela School of Medicine, UKZN
719 Umbilo Road, Durban, South Africa.
Director: Prof. Tulio de Oliveira





