When Lineages Reappear
text: Graeme Dor

Remember when Omicron was first detected in South Africa in late 2021?
Three lineages emerged almost simultaneously: BA.1, BA.2, and BA.3. While BA.1 drove the initial global wave, it was BA.2 that persisted and ultimately gave rise to most of the lineages circulating today. BA.3, in contrast, appeared to fade. After mid-2022, it was no longer detected, and was largely considered an evolutionary dead-end.
More than two years later, in November 2024, a highly divergent descendant of BA.3, designated BA.3.2, was identified in South Africa. What makes this lineage unusual is not just its reappearance, but how it re-emerged, on a long evolutionary branch, with no intermediate genomes detected. This suggests a prolonged period of undetected evolution, followed by sudden re-entry into the sampled population.
This matters because it highlights a pathway of viral evolution that is difficult to observe in real-time. Rather than accumulating changes gradually under continuous surveillance, lineages like BA.3.2 may evolve in relative isolation, such as in chronically infected individuals, before re-emerging with substantial genetic differences.
Despite low circulation through much of 2025, BA.3.2 spread globally and began increasing in multiple regions towards the end of the year. By early 2026, it had been detected across multiple continents, reaching notable proportions in some countries (Figure 1) . Genetically, it carries extensive changes in the spike protein, including key mutations (Figure 2) associated with immune escape and receptor binding, raising the possibility of altered biological properties that could enhance its spread. In many settings, it has also been observed more frequently in children, potentially reflecting interactions between its mutational profile and existing patterns of population immunity.
For public health, the key insight is not just the emergence of BA.3.2 itself, but what it represents. Even as dominant SARS-CoV-2 lineages continue to circulate globally, divergent variants can evolve in parallel and reappear unexpectedly.
This discovery also reinforces the critical role of genomic surveillance in southern Africa. The detection of BA.3.2 reflects sustained sequencing efforts in the region, which remain essential for identifying and characterising emerging variants before they spread more widely.
As SARS-CoV-2 continues to evolve, BA.3.2 highlights an important ongoing risk, that seemingly extinct lineages may not be gone, but instead evolving out of sight, with the potential to re-emerge under the right conditions.
For acces to the full publication, visit: https://doi.org/10.1093/ve/veag016
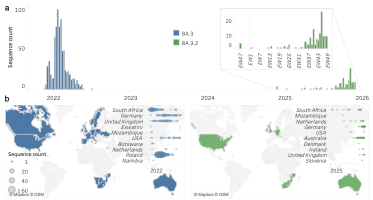
FIGURE 1: Weekly spatio-temporal genomic distribution of BA.3 lineages globally. (a) Time-series distribution of BA.3 sequences by epidemiological week (EW), stratified by Pango lineage (BA.3 shown in blue, BA.3.2 shown in green). The inset provides an expanded view beginning from the first detection of BA.3.2 in November 2024. (b) Geographic distribution of BA.3 (blue) and BA.3.2 (green) detections, with dot plots reflecting weekly sequence counts for each country. Countries are ordered by the date of first detection, and limited to the ten earliest-reporting countries.
News date: 2026-05-13
Links:
KRISP has been created by the coordinated effort of the University of KwaZulu-Natal (UKZN), the Technology Innovation Agency (TIA) and the South African Medical Research Countil (SAMRC).
Location: K-RITH Tower Building
Nelson R Mandela School of Medicine, UKZN
719 Umbilo Road, Durban, South Africa.
Director: Prof. Tulio de Oliveira




